Convert Oracle XMLTYPE to Oracle tables
In this example we will read XML data from a table with an XMLTYPE column in the Oracle database and convert the XML to tables in the same Oracle database.
We will use Flexter to extract the data to and from the same database in this example.
We need to give Flexter table and column name that contains the XML data.
JDBC input sources require us to include the input path with a jdbc: protocol.
[flexter_banner]
|
1 2 3 |
$ xml2er -T <[[Catalog.]Schema.]Table]> -C <XML Column> -U <user> -P <password> [OPTIONS] jdbc:<database url> # or using a sql query $ xml2er -T "<SQL QUERY>" -C <XML Column> -U <user> -P <password> [OPTIONS] jdbc:<database url> |
Example:
|
1 2 3 |
$ xml2er -T tab -C xml_col -U rob -P rob_pass [OPTIONS] jdbc:oracle:thin:@//d.org.io:<port-no> # or using a sql query $ xml2er -T "select xml_col from tab" -C xml_col -U rob -P rob_pass [OPTIONS] jdbc:oracle:thin:@//d.org.io:<port-no> |
This command will create one connection and against the source database and retrieve the data sequentially.
To read in parallel with multiple threads (available on a single machine as well), a sequential ID column name and its range must be provided. The options which we are using are given in the below table
|
1 2 3 4 5 |
-T, --table TABLE Table or SQL which brings XML data -C, --column COLUMN Column which brings XML data -I, --part-in-col PART_COL Partition or primary key column -<MIN>, --part-in-min MIN Lower bound to retrieve rows +<MAX>, --part-in-max MAX Upper bound to retrieve rows |
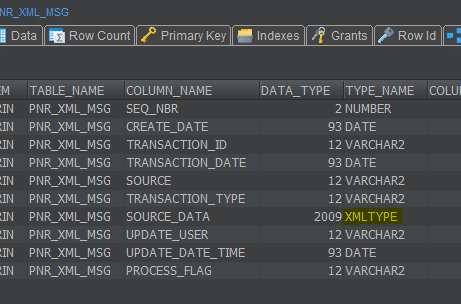
We have a table “pnr_xml_msg” which has xml content in the column “source_data”. The Primary Key is seq_nbr. Let’s take a look at the table and the xml content that will parse with Flexter.
We can see that the table contains the “source_data” column of XMLTYPE. This column contains the xml data.
[flexter_button]
We then try to generate a logical schema based on the xml content in the column
|
1 2 3 4 5 6 7 8 |
# running the command with -s option $ xml2er -s \ -T "select source_data,seq_nbr from pnr_xml_msg where seq_nbr between 495301 and 495321" \ -C source_data \ -I seq_nbr -495301 +495321 \ -U <source username> \ -P <source password> \ jdbc:oracle:thin:@//d.org.io:<port-no> |
If the flexter command works fine, we can proceed with generating the logical schema for the xml content in the table
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 |
# generating the logical schema for the xml content $ xml2er -g3 \ -T "select source_data,seq_nbr from pnr_xml_msg where seq_nbr between 495301 and 495321" \ -C source_data \ -I seq_nbr -495301 +495321 \ -U <source username> \ -P <source password> \ jdbc:oracle:thin:@//d.org.io:<port-no> \ # schema origin: 22 logical: 7 job: 34 # statistics startup: 3699 ms parse: 4633 ms stats: 6065 ms map: 1677 ms unique xpaths: 227 |
Here, we are using -U and -P parameter as the input username and password to the jdbc URI
|
1 2 3 |
-U, --iuser USER Input user -P, --ipassword PASSWORD Input password -F, --input-format FORMAT Input format. (jdbc,parquet,orc,json,csv, tsv) |
Now as we have generated the logical schema we can extract the data to the output jdbc connection. We can use the following options to generate the output as desired
|
1 2 3 4 5 6 7 8 9 10 11 |
-o, --out OUTPUT Output location path -u, --ouser USER Output user -p, --opassword PASSWORD Output password -B, --batchsize BATCHSIZE Batch size to write into databases default: 1000 -f, --format FORMAT Output format. (jdbc, parquet, orc, json, csv, tsv) -z, --compression COMPRESSION Parquet, csv, tsv compression mode (none, snappy, gzip, lzo, lz4, bzip2, xz) -S, --savemode SAVEMODE Save Mode when table, directory or file exists ex: [e]rror, [a]ppend, [o]verwrite, [i]gnore default: append |
Initially we use the the primary key to partition the data we are extracting using the -I option, We can also parse and extract the entire XML content if we skip the -I option
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 |
$xml2er -l7 \ -T "select source_data,seq_nbr from pnr_xml_msg where seq_nbr between 495301 and 495321" \ -C source_data \ -U <source username> \ -P <source password> \ jdbc:oracle:thin:@//d.org.io:<port-no> \ -o jdbc:oracle:thin:@//d.org.io:<port-no> \ -u <target username> \ -p <target password> # output path: jdbc:oracle:thin:@//d.org.io:<port-no> user: <target username> password: *** format: jdbc # schema origin: 22 logical: 7 job: 35 ... 16:37:10.426 INFO table EMD_Info: creating 16:37:11.337 INFO table EMD_Info: writing 16:37:14.849 INFO table Ticketing: creating 16:37:15.144 INFO table Ticketing: writing ... # statistics startup: 4097 ms load: 9211 ms parse: 1693 ms write: 54730 ms stats: 1526 ms map: 4 ms unique xpaths: 227 |
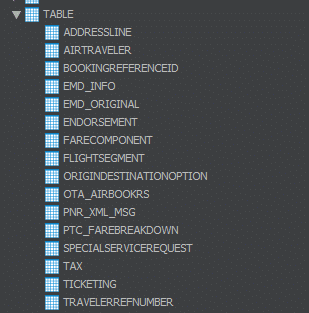
In the above output we can see that the Flexter is writing the tables in the target db as shown in the output. The XML content parsed can be seen in the form of tables in the given jdbc path.
We can see that the parsed tables are created in the target db as per the jdbc path.
We can run basic queries to see if the data is parsed correctly
|
1 2 3 4 5 6 7 8 9 10 11 12 |
sql> describe tax; sql> describe ticketing; sql> describe airtraveler; sql> select * from tax limit 5; FK_PTC_FAREBREAKDOWN AMOUNT CURRENCYCODE TAXCODE 0000000000000000082 0.91 EUR EX 0000000000000000012 6.5 EUR HB 0000000000000000006 21.06 CHF UP 0000000000000000012 16.04 GBP UB ... sql> select * from <table-name> where <condition> ... |
[faq_button]




